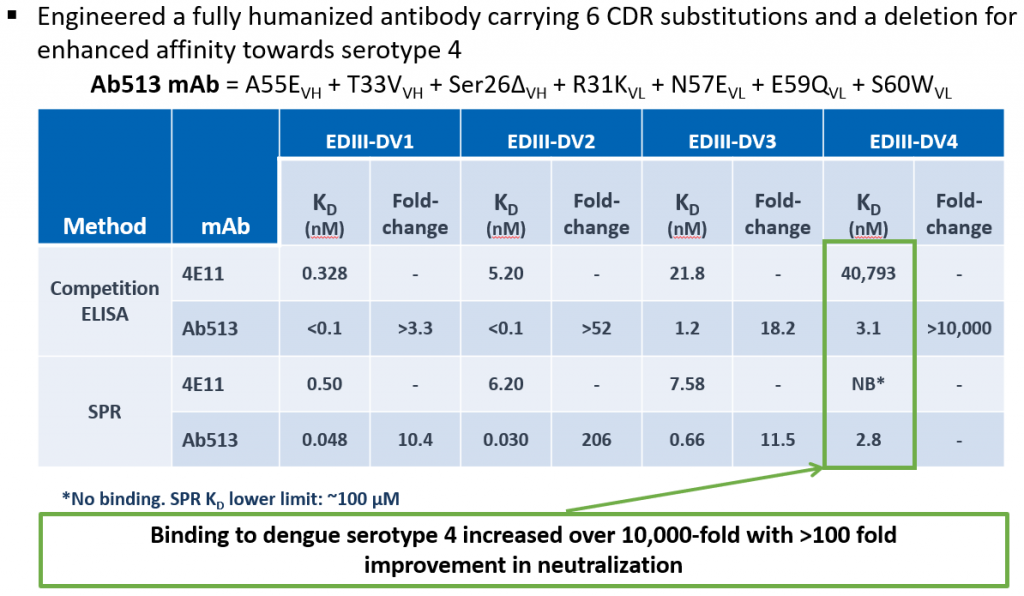
The 4E5A antibody that was obtained by redesigning 4E11 showed a significantly higher affinity to DENV-4 strains compared to 4E11. However, its binding affinity for different strains of DENV-4 was still weaker relative to that for DENV-1, -2, and -3 serotypes. To enhance further enhance the binding affinity of 4E5A to DENV-4 while retaining the strong binding affinity to the other serotypes, we further developed our tools to incorporate a network-based analysis using graph theory approaches.
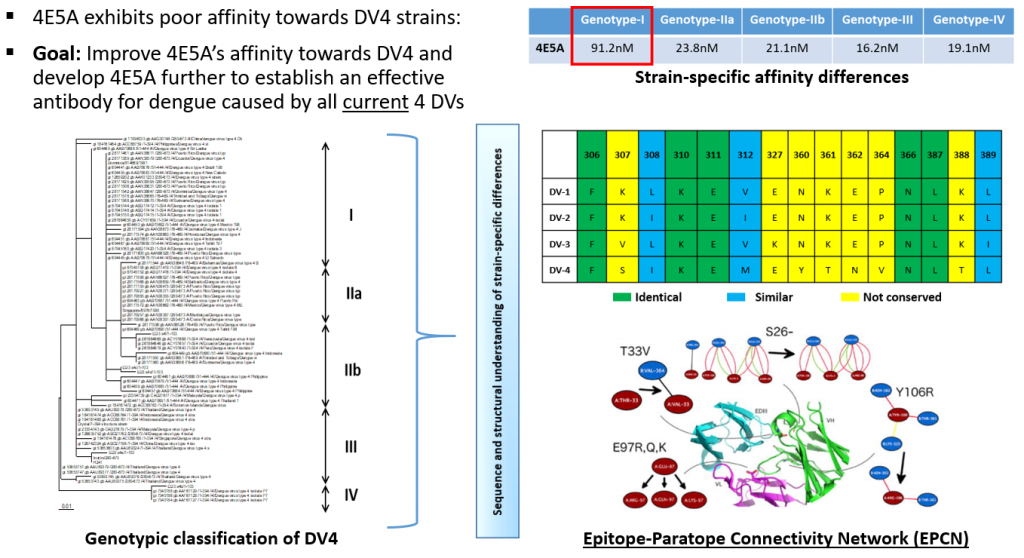
Both short-range and long-range interactions between non-bonded residues at the epitope-paratope interface were represented as a two-dimensional map (epitope paratope connectivity network or EPCN) and each residue was scored based on its connectivity to other residues in this interface. This network representation permitted investigating in detail the impact of amino acid changes such as mutations, residue deletions, and residue insertions in altering the associated networks. This approach was different from just viewing amino acid changes as conservative and non-conservative changes respectively as even conservative substitutions substantially impacted the connectivity of the residues in the network. Using this network-based approach, the 4E5A antibody was further redesigned by introducing two mutations in HCDR1, which included a single residue deletion, that improved the EPCN and enhanced its affinity to all DENV serotypes, and to make it clinically developable as a potent pan-DENV therapeutic. The overall affinity gain of the engineered antibody was 13,000-fold for DENV-4, which ranks amongst the largest affinity enhancement reported by in silico antibody affinity enhancement studies.