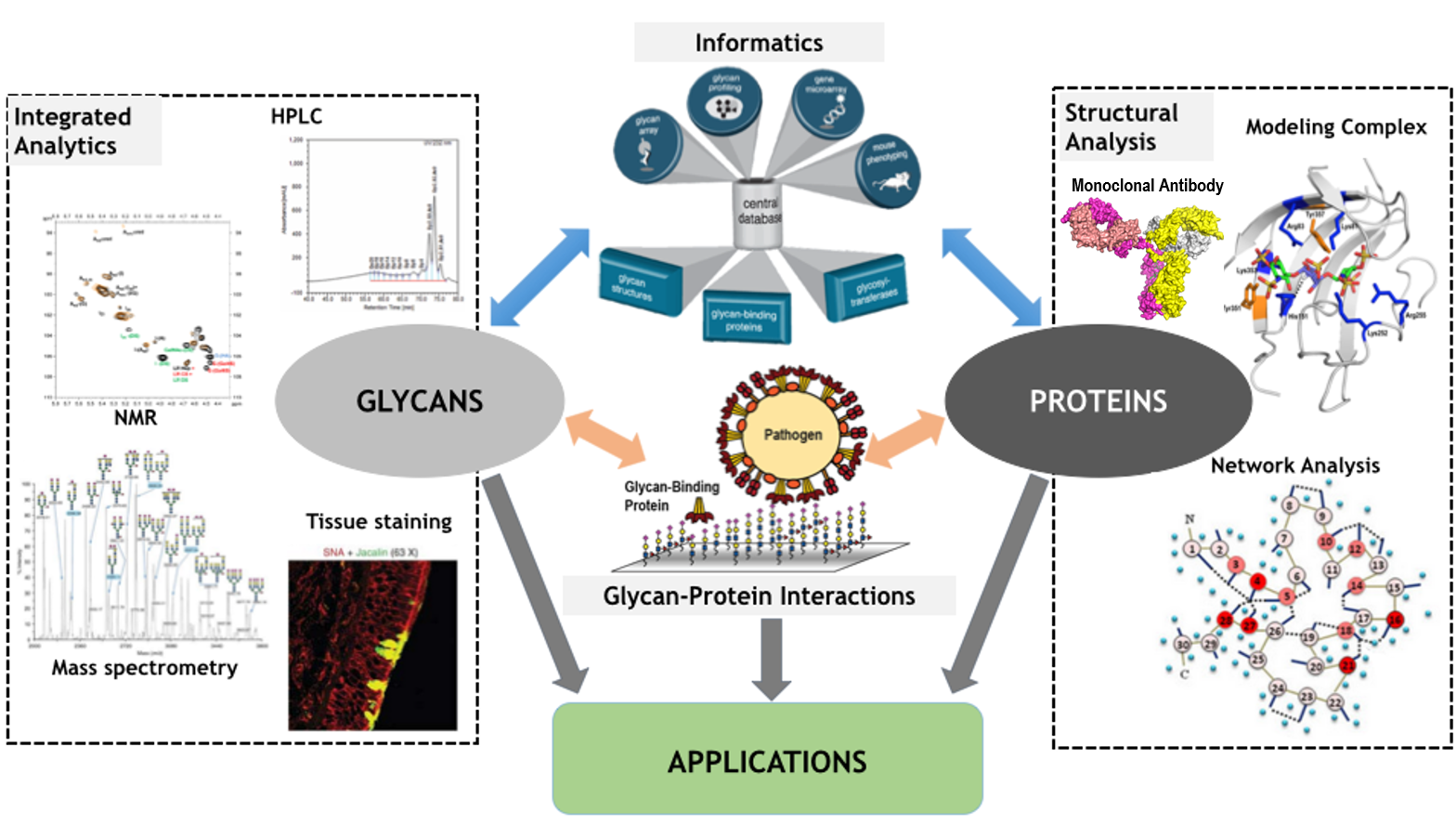
The Sasisekharan Lab rests on a foundation of diverse glycomics technologies developed and applied over the years. We have tackled underlying problems in glycomics and focus on the fundamental aspects of glycan-protein interactions. Bringing various approaches to bear on these problems has been a principal area of focus for the group.
We study glycan structure (both glycosaminoglycans (GAGs) and branched glycans) through the use of orthogonal analytical approaches (NMR, MS, enzymology etc.,). In a simultaneous and complementary effort, we developed methods to describe protein-glycan interactions using structure-based tools developed by us and experimental/heuristic approaches such as through conformational studies using NMR and our informatics platform. Finally, we directly probe essential glycan-protein interactions, such as specificity and affinity, utilizing a novel set of array and tissue-based studies.
Our lab originated our efforts with the GAG heparin. That work developed the needed analytical and enzymatic tools with defined substrate-product profiles to understand their structure-function relationships. We also set out to create distinct and novel tools to study glycans, tools which are uniquely suited to their chemical diversity and conformational plurality in the context of their protein-binding partner. These tools became core to our ability to investigate different glycan-protein model systems. To this end, as mentioned above, we developed a graph theory/network/informatics-based approach to investigate protein structure, especially in the context of glycan interactions.
It has become ever more evident to us that to understand glycan-protein interactions as a whole – given the ‘analog’ continuum of their interaction features – that we need to capture diverse sets of interactions to help us refine our strategy for integrated (glycan and protein-based) ‘glycomics tool’ building.
Our work in the glycan-protein interactions area led us to the infectious diseases arena, focusing on Influenza, flaviviruses (such as as Dengue and Zika). It also led to developing antibody engineering methods by incorporating network-based analysis (using graph theory) as outlined in the Antibody Engineering Strategy section.
Antibody engineering, taken together with integrated analytics, has enabled us to develop a Rapid Response Regulatory Framework for emerging infectious diseases. This framework has led to biologics development (including GMP manufacturing and non-clinical studies) and first-in-human studies with unprecedented timelines to tackle the Zika virus (in nine months), Yellow Fever virus (in about six months), and SARS 2 (COVID19) virus (in about four months).
Glycan Tools Related Areas
Glycan-Protein Structure-Function Relationship
