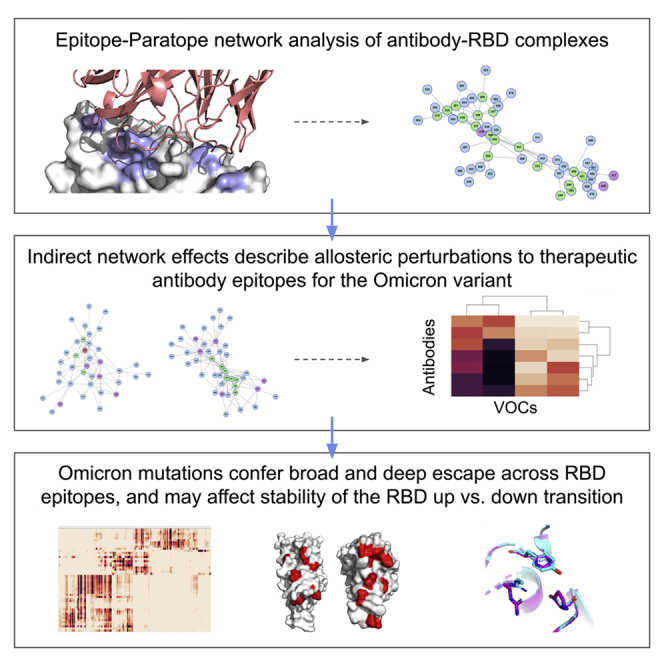
In October-November 2021, after the successful rollout of vaccines to counter the deadly Delta variant of SARS-COV-2 global outbreak, a new threat was looming. There were reports of a new variant of SARS-COV-2 known as Omicron, which was wreaking havoc globally given that it carried a large number of mutations that were likely to render previous exposure to older variants, or vaccination useless in providing protection against this new variant. In a similar manner, there was a growing concern about this variant in rendering the approved therapeutic monoclonal antibody treatments against SARS-COV-2 ineffective.
Companies such as Regeneron Pharmaceuticals put out a press release that their approved antibody cocktail shows lower neutralization based on detailed structural modeling and preliminary in vitro data, other companies such as Adagio Therapeutics based on just sequence conservation of the epitope stated that their lead candidate ADG20’s neutralization potency would not be impacted by Omicron.
Sasisekharan Lab had developed several tools to investigate the mutational constraints on epitope surfaces going beyond sequence conservation to include analysis of the inter-residue interaction network of a residue. Therefore, Sasisekharan Lab set out to map the mutational landscape of SARS-COV-2 in the context of the emergence of the Omicron variant using network-based analyses tools the Lab developed.
Sasisekharan Lab mapped the epitope surface on the receptor-binding domain (RBD) of SARS-COV-2 using amino acid interaction (AAI) networks, which are well suited for interrogating constellations of mutations that function in an epistatic manner. Using AAI, Sasisekharan Lab mapped the Omicron mutations directly and indirectly driving increased breadth and depth of escape from vaccines and therapeutic mAbs. Further, Sasisekharan Lab analyzed the epitope networks for authorized therapeutic antibodies and assessed perturbations to each antibody’s epitope.
Sasisekharan Lab predictions were realized by experimental findings of Omicron neutralization escape from therapeutic antibodies ADG20, AZD8895, and AZD1061. Importantly, the AAI predicted escape resulting from indirect epitope perturbations was not captured by previous sequence or point mutation analyses. Specifically it predicted that ADG20’s potency would be impacted by the Omicron mutants through indirect network perturbations and this finding was borne out by the subsequent neutralization studies which showed that ADG20 activity dropped by 300 fold.
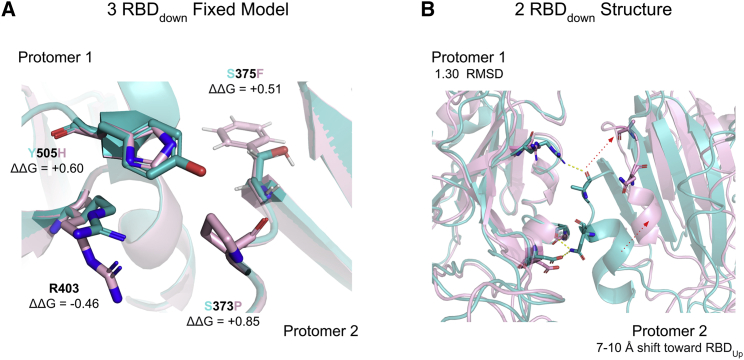
The effects of Omicron mutations on the RBDdown-RBDdown interface
Finally, for several Omicron RBD mutations, Sasisekharan Lab found evidence for a plausible role in enhanced transmissibility via disruption of RBD-down conformational stability at the RBDdown-RBDdown interface.