Evolution of Antibody Engineering Platform in the Sasisekharan Lab
Since 2010, the Sasisekharan Lab has been systematically building tools for computational antibody engineering with the support of over two dozen researchers (undergraduates, graduate, and post-doctoral students, as well as Research Scientists), and two significant NIH-funded grant awards (including an NIGMS merit award) and Agilent Thought Leader Award. This also led to the creation of commercial opportunities which included technology licensing as well as the creation of the biotechnology companies Visterra (which was acquired by Otsuka Pharmaceutical for 430 million USD) and Tychan Pte Ltd. in Singapore.
A common theme connecting our scientific and engineering journey is a focus on the optimization of interfaces between antibodies and defined target epitopes with the goal of generating novel antibodies with improved properties that are clinically developable. Additionally, we have continued to develop the platform with significant improvements resulting in the delivery of several clinical-stage antibodies.
The platform’s success began with the ‘Epitope’ (i.e.. the target) focused antibody affinity enhancement, and has arrived at ‘Epitope agnostic’ affinity enhancement of antibodies. Over the last decade the mAb PDB has grown from 1,700 structures to 7,000 and mAb sequences have grown from 50,000 to over a billion. The explosion in structure and sequence data makes this platform very useful and hence impactful.
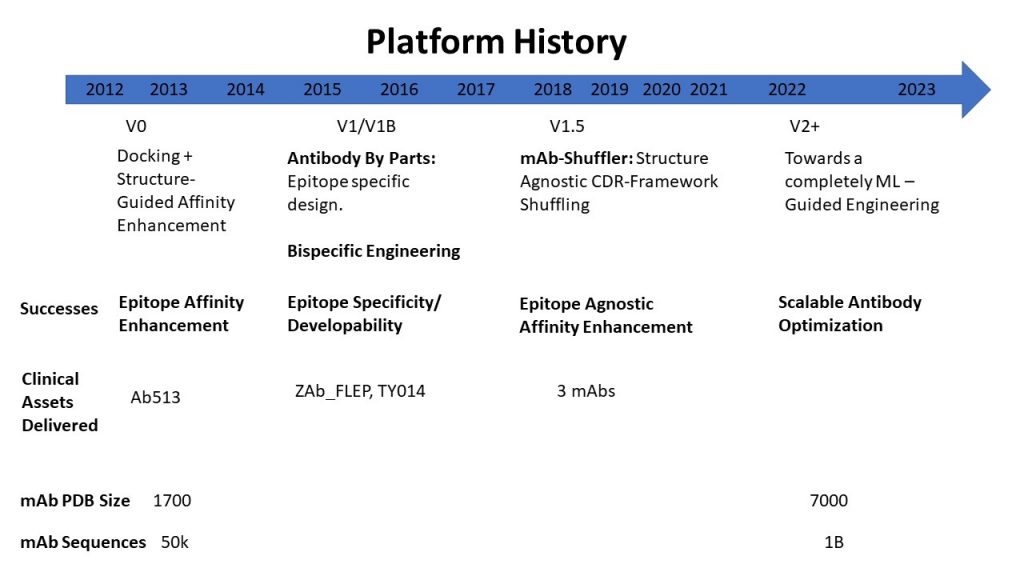
Sasisekharan Lab’s successful journey in Antibody Engineering
The V0 Platform
During the 2012-13 time frame, the Sasisekharan Lab introduced the concept of ‘featurization’ of amino acid properties and antibody–antigen interfaces. Today such featurization methods are widely used in machine learning (ML) guided protein engineering. ‘Featurization’ is capturing probable attributes of amino acids which are used in models to predict functional attributes.
For example, by using amino acid featurization, the Sasisekharan lab successfully trained a multivariate logistic regression (MLR) model (which broadly represents ML in the present day) to discriminate native from decoy antigen-antibody complexes. This was challenging at the time due to limited structural and sequence information as noted earlier.
As a clear demonstration of success associated with these early tools and methods (Referred to as V0 platform), Sasisekharan Lab reliably predicted antibody-antigen interactions. This enabled the engineering of an antibody to increase its affinity towards a specific Dengue virus-4 envelope protein antigen over 450 fold, while modestly improving its affinity to 3 other antigens viz. DENV-1 -> DENV-3 (see Dengue story)
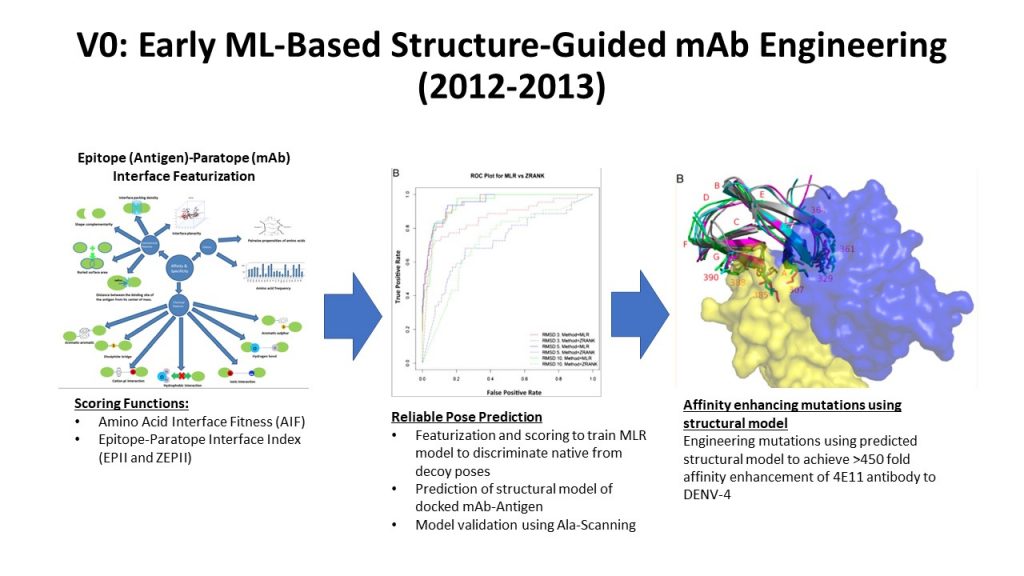
Early Machine-Learning Guided Platform For Antibody Affinity Enhancement
Accomplishments of the V0 Platform: >450-fold affinity enhancement to DENV-4 and retaining (modest improvement) to the affinity for DENV-1-3 with less than 150 distinct antibody constructs screened when no crystal structure of 4E11 was available. This kind of affinity enhancement by screening so few constructs could not be accomplished by previous structure-guided and phage display methods. This was a tour-de-force accomplishment that was not possible then and it led to a potent therapeutic candidate (Ab513) that was validated in pre-clinical in vivo studies and Phase 2 human clinical studies are currently underway.
Although the featurization tools were used to optimize a specific antibody to improve its affinity, the potential utility of these tools goes well beyond single antibody optimization.
The V1 Platform (3DMAbDesign)
During the 2014-17 timeframe the Sasisekharan lab expanded these tools, adding the ability to analyze interatomic amino acid networks. This enabled
- identification of optimal target epitopes that were most mutation-resistant due to structural constraints and
- engineering antibodies by combining complementarity determining regions or CDRs and variable domains from various antibody data sources.
This methodology led to the generation of target-specific antibody libraries. In other words, we could assemble parts CDRs and framework or FW regions from different antibodies to create a novel antibody which optimized contact with the target epitope.
This enabled the Sasisekharan Lab to create an antibody engineering platform (3DMAbDesign or the V1 platform) that assembled known antibody parts using various features and scoring functions that the Sasisekharan Lab had developed as a part of the V0 platform to rapidly create new antibodies with distinct functional properties for emerging infectious diseases.
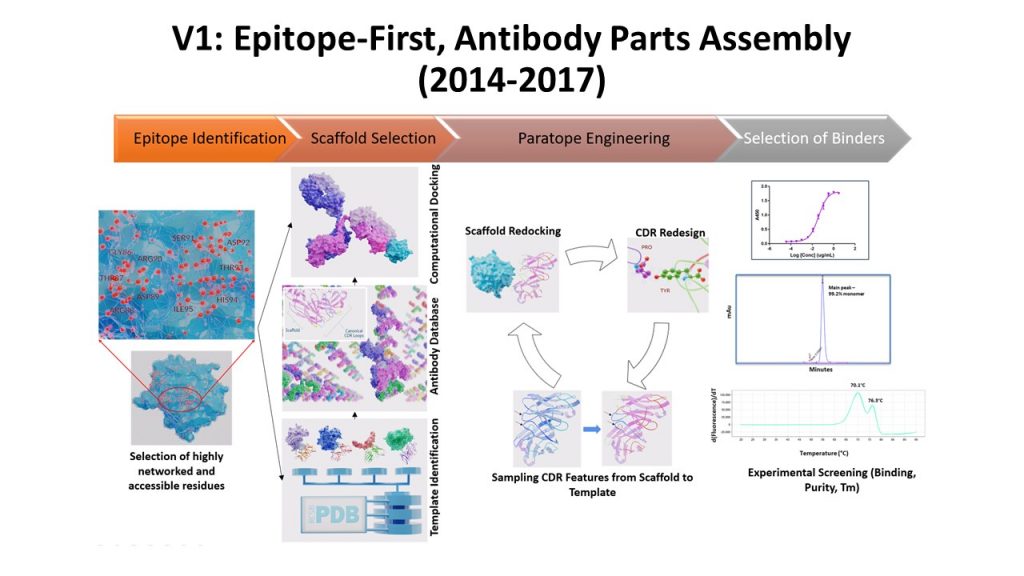
3DMAbDesign Platform: Engineering Antibodies by Assembling Parts
Accomplishments of the V1 Platform: The parts assembly approach explained in Antibody Engineering Methods and illustrated in an animation enabled rapid engineering of antibodies that specifically target a defined epitope. This platform made it possible to develop therapeutic antibodies against the ZIKV in 2017 and the Yellow fever virus in 2019. The engineering platform together with the regulatory framework for comprehensive integrated-analytics-driven CMC and non-clinical studies permitted rapid completion of IND-enabling studies within nine months for the Zika-virus antibody and seven months for the Yellow fever antibody.
The Sasisekharan Lab focused on this challenge with support from Agilent (through their Thought Leader award) and the Labs’ participation in Singapore-MIT Alliance for Research and Technology (SMART, Singapore). Singapore was significantly affected by SARS in 2003, and the Ebola outbreak in 2014, which highlighted the need to address how to respond in a time frame of months, rather than years, to create new antibody therapeutics.
The Bispecific mAb V.1B Platform:
Not only did the Sasisekharan Lab advance engineering of monospecific antibodies, but the Lab also came up with unique approaches to design bispecific antibodies (Platform V1B, 2015-19). By building on the interface featurization aspects of the V0 platform to engineer mutations in the CH1-CL and CH3-CH3 interfaces Sasisekharan Lab generated IgG-like bispecific antibodies using a single cell culture system with standard antibody purification methods.
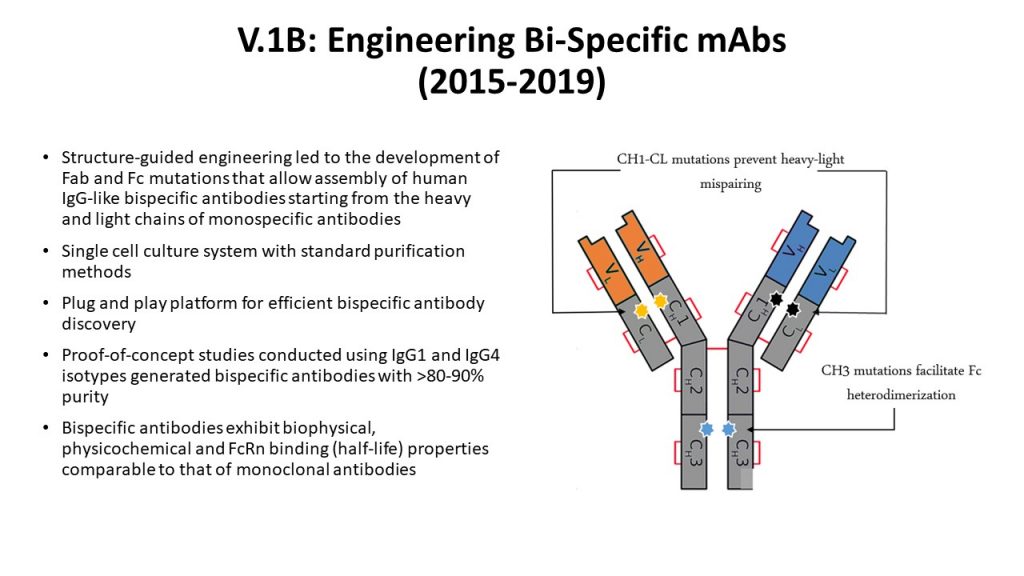
Bispecific Antibody Engineering Platform
Accomplishments of the Bispecific V.1B Platform: The rationally engineered mutations focusing on the constant domain interfaces (CH1-CL and CH3-CH3) readily permitted the generation of bispecific antibodies of high purity (>80%) from a single transient transfection. Since the mutations were made at sites that are evolutionarily conserved across Ig genes, they permit BsAb assembly from antibodies belonging to diverse germline origins and isotypes.
Journey through the COVID-19 pandemic and beyond
The Sasisekharan Lab pursued the advancement of the antibody engineering platform during challenging times. We built on the antibody parts assembly to develop a platform that shuffles the FR and engineered CDR regions selected from millions of antibody sequences obtained from next-generation sequencing methods. This shuffling approach permitted optimization of antibodies selected from germlines that are enriched upon infection by pathogens such as SARS-COV-2 to enhance their affinity by 7-10 fold in a rapid manner with an experimental screen size of 80-100 designs.
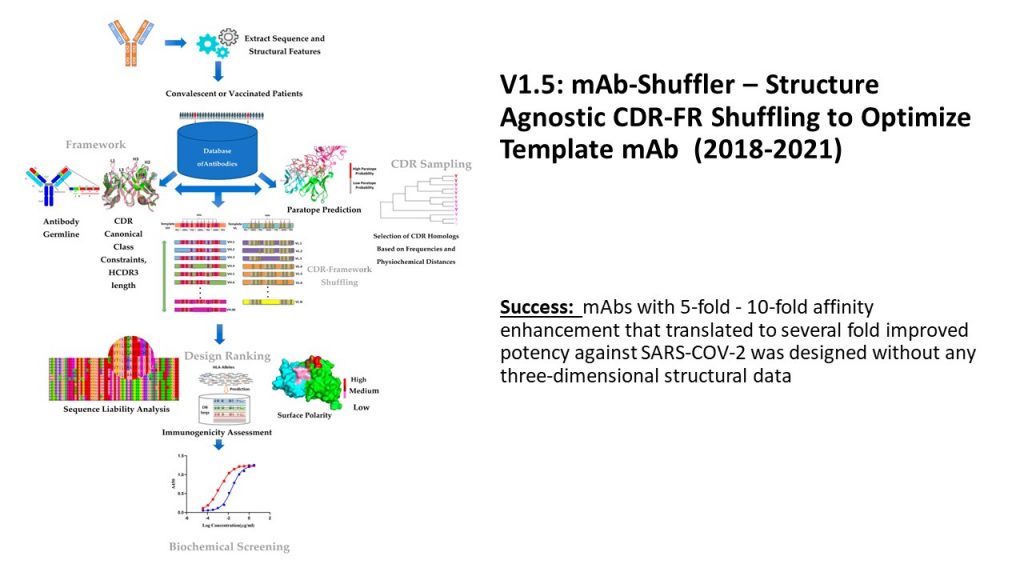
Shuffling CDR-Framework Regions for MAb Affinity Enhancement
Accomplishments of the V1.5 Platform: This platform builds on the V1 Platform that was used to generate the anti- Zika virus antibody by expanding the search for CDRs and scaffold regions to the millions of human antibody sequences generated through next-generation sequencing. Based on this expanded search of a much larger antibody sequence space, the Sasisekharan Lab was able to shuffle and assemble CDR and FR parts, without any guidance from the X-ray co-crystal structures of antibodies or antigens, to engineer antibodies with improved affinity. This approach was adapted to engineer a SARS-COV-2 antibody with improved affinity and potency relative to the starting template antibody that was initially used for searching FR and CDR sequence space.
Towards implementing ML methods for antibody engineering
With the success of Alphafold which is a deep-learning neural network that accurately predicts the three-dimensional structure of proteins, ML tools are gaining prominence for their use in drug design, particularly in the context of antibody engineering. The Sasisekharan Lab has stayed on track in developing more advanced tools in the context of mapping epitope surfaces and improving the featurization of antibody-antigen interfaces to make them amenable for use with the current ML models and tools.
Network/topological maps of epitope surfaces: One area of study was uncovering novel patterns in glycoepitopes to enable complex topological search of similar glycoepitopes across pathogens. With this understanding, we could rapidly repurpose antibodies targeting these patterns against a range of pathogens. The other area involved a network-based approach to mapping the entire epitope surface of antigens that allowed accurate prediction of how antigenic evolution of SARS-COV-2 impacted the potency of therapeutic antibodies such as Adagio’s ADG-20.
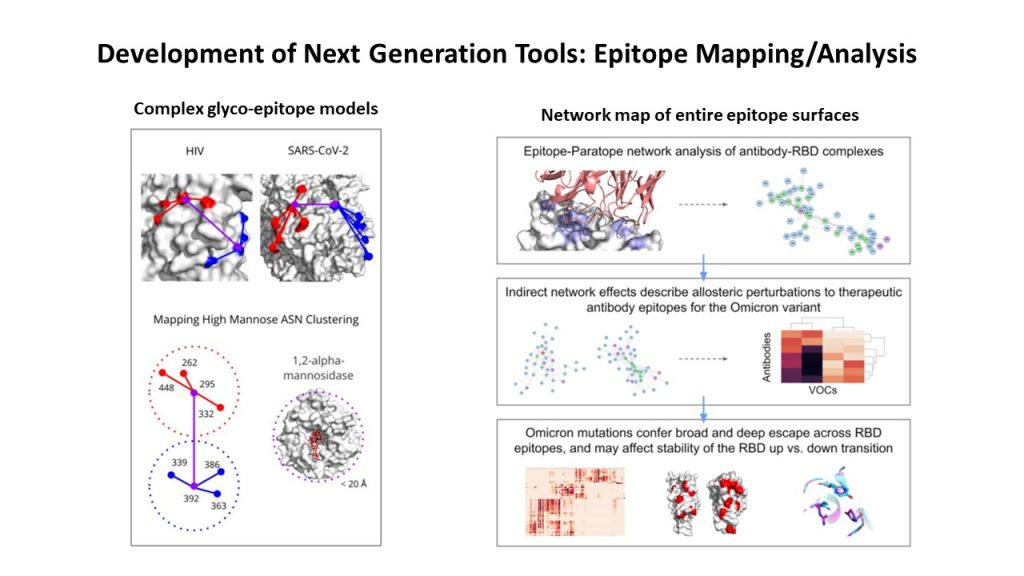
Complex epitope mapping tools for rapid antibody engineering
Practical applications of network-based epitope mapping: In late November 2021, Adagio Therapeutics asserted that their antibody ADG-20 would neutralize the Omicron variant of SARS-COV-2 which had already escaped neutralization from vaccines and other clinically approved antibodies at that time. They came to this conclusion based on sequence similarity in the epitope surface (targeted by ADG-20) between the Omicron variant and the past variants. This belief assumed that epitope regions are similar if there is substantial overlap between the sequences of the epitope regions of different viral strains. However, this belief misses important nuances to the perturbations caused by amino acid changes to the entire inter-residue interaction network involving other residues that are directly within the epitope surface and just outside it, and how these perturbations impact the binding of an antibody to the epitope surface.
In early December 2021, the Sasisekharan Lab used network methods to predict that the epitope surface changes on Omicron would reduce the potency of many of these SARS-COV-2 antibodies including ADG-20. A week later it was reported that ADG-20 took a 300-fold hit in neutralization potency against Omicron This validated the predictions from the Sasisekharan lab and highlighted the limitations of sequence analyses.
Augmented learned features for predicting antibody-antigen binding affinity :Building on Sasisekharan Labs’ work done in 2013 on featurizing antibody-antigen interfaces with limited structural data, the Lab expanded the featurization to learn from much larger structural and sequence datasets of antibody-antigen complexes annotated with binding affinity information to classify features that correlate with high vs. low binding affinity. These features are amenable to developing and implementing various ML algorithms to rapidly optimize antibodies in a scalable fashion.
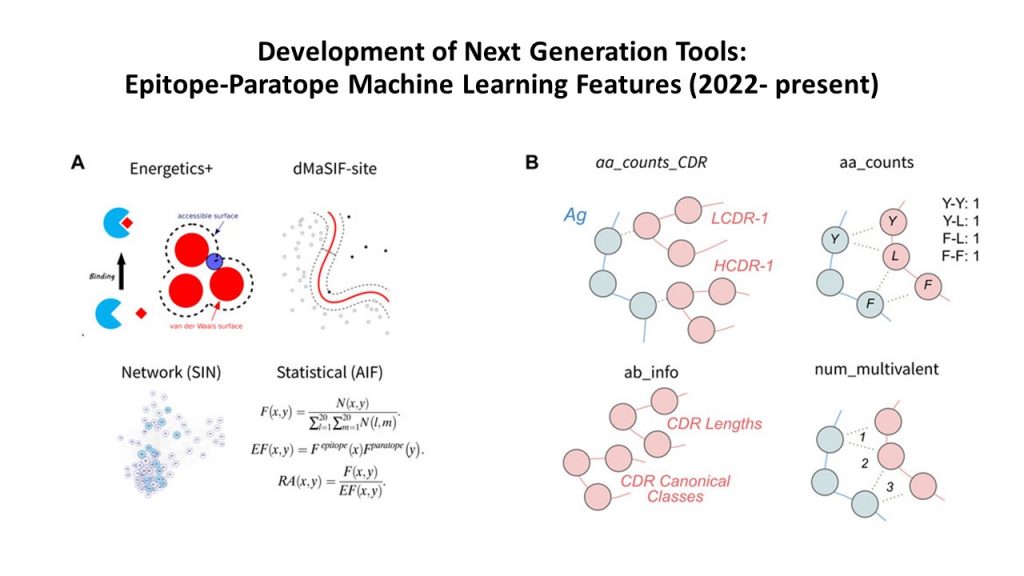
Learned features discriminate antibodies with low and high antigen-binding affinity
Summary of the Journey and Future Directions
The Sasisekharan lab has had a successful journey in the field of antibody engineering with an overarching goal of providing rapid, accessible, and affordable therapeutic solutions against global health threats including the most recent COVID-19 pandemic. Specifically, the Sasisekharan Lab’s approach can rapidly generate solutions by taking advantage of prior knowledge of a biologically relevant target. Studies describing approaches similar to assembly of known parts to engineer antibodies have been published recently, highlighting the increasing value of this approach.
The vision of the lab even back in early 2010’s was consistent with the current thinking of using ML models to design better antibodies by taking advantage of the wealth of known information and datasets. These approaches have challenged the view that it is implausible for closely related polypeptide sequences to have substantially diverse functional properties. In other words, sequence similarity is considered the most deterministic of functions – this doesn’t always hold. With the rapidly growing NGS datasets, we now know that diverse VH and VL genes can be used to produce antibodies against a particular protein epitope; likewise, closely related antibodies can have different target specificities.
With the increasing application of ML-based tools for antibody design and engineering technology, there are two categories where such tools are being implemented:
- to design libraries in the order of 10^6-10^10 to pan and identify antibodies optimized for various properties and;
- to optimize antibodies by generating 10s of 1000s of designs for experimental screening.
The Sasisekharan lab’s vision for implementing ML in any therapeutic antibody design and engineering problem stems from an intelligent sampling of allowed mutations (classified based on ML models that use our featurization strategies) to enhance antibody binding affinity which would permit reliable antibody optimization in weeks involving less than 200 designs.
